Bioinformatic analysis
1 March 2024
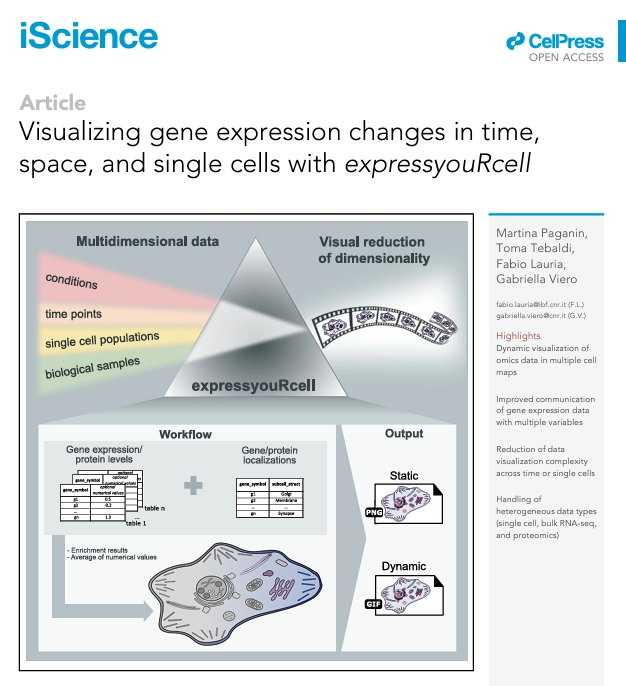
Visualizing gene expression changes in time, space, and single cells with expressyouRcell
Paganin M, Tebaldi T, Lauria F, Viero G. Visualizing gene expression changes in time, space, and single cells with expressyouRcell. iScience. 2023 May 11;26(6):106853. doi: 10.1016/j.isci.2023.106853. PMID: 37250782; PMCID: PMC10220493.
The last decade has witnessed massive advancements in high-throughput techniques capable of producing increasingly complex gene expression datasets across time and space and at the resolution of single cells. Yet, the large volume of big data available and the complexity of experimental designs hamper an easy understanding and effective communication of the results. We present expressyouRcell, an easy-to-use R package to map the multi-dimensional variations of transcript and protein levels in dynamic cell pictographs. expressyouRcell visualizes gene expression variations as pictographic representations of cell-type thematic maps. expressyouRcell visually reduces the complexity of displaying gene expression and protein level changes across multiple measurements (time points or single-cell trajectories) by generating dynamic representations of cellular pictographs. We applied expressyouRcell to single cell, bulk RNA sequencing (RNA-seq), and proteomics datasets, demonstrating its flexibility and usability in the visualization of complex variations in gene expression. Our approach improves the standard quantitative interpretation and communication of relevant results.
Keywords: Computational bioinformatics.

