Blog
13 April 2026

What is Translatomics? The Missing Layer in Gene Expression
Many
molecular biologists have reported that RNA-seq data show upregulation of a key
gene, but protein levels do not match. Or the crucial transcriptional hits are
completely absent from proteomics data, highlighting the “disconnect” between
transcriptomics and proteomics data.
In this blog article, we summarise the measurable factors that affect translation, and
why it is becoming increasingly important to look beyond conventional molecular
methodologies to get the bigger picture. But first, let us understand the
biological mechanisms that influence the translation process.
Why mRNA ≠ protein: the biology behind the “disconnect.”
Vogel and Marcotte, quoted in their 2012 article, “Current data demonstrate a substantial role for regulatory processes occurring after mRNA is made”. Many researchers have pointed out that after transcriptomics, cells undergo a decision process of what should be translated and how much. One pan-cancer proteogenomic study has confirmed the disconnect between mRNA and protein by analyzing more than 2,555 tumor samples across 15 cancer types, revealing that the correlation coefficient between transcriptomic and proteomic measurements typically ranges from 0.3 to 0.5 [1].
This implies that researchers might be:
• Focusing on transcriptional changes that do not lead to functional proteins
• Overlooking crucial regulatory mechanisms that operate between RNA and protein;
• Drawing conclusions about biological processes without enough data
To understand various
cellular processes and disease etiology, it has become necessary to measure the
biological mechanisms that explain the correlation between mRNA abundance and
protein variation. Key factors that help in decoding the disconnect are:
- Translation efficiency – Some mRNAs are translated frequently, while
others remain underutilized despite being abundant. A highly abundant
transcript with low translation efficiency produces far less protein than
a rare transcript that's being translated intensively.
- Ribosome occupancy – Actively translated transcripts may be
loaded with many ribosomes at once (polysomes), while repressed mRNAs
usually have few or none. This explains why two transcripts with similar
abundance can have different protein outputs.
- Translational control: Cells can reprogram translation within
minutes in response to stress, signalling, or developmental cues. These
rapid changes happen long before mRNA levels shift, which typically takes
hours.
Translation regulation thus alone exerts greater control over protein output than transcription, mRNA degradation, and protein degradation combined. Thus, molecular biologists are moving towards the “Translatomics” approach that lies between transcriptomics and proteomics. Rather than measuring what mRNAs are present, translatomics tells us which mRNAs are actually being translated into proteins at any given moment, helping explain the missing 60–70% of protein variation [1].
Decoding translatomics: from ribosomes to tRNAs
The
techniques for studying translation have expanded considerably over the past
decade. Each approach captures a different layer of the process, from tracking
ribosome-embedded mRNA to measuring tRNA modification and decoding translation
efficiency. Understanding what these tools measure and where they differ is
useful for deciding which approach fits a given research question. The core
methods are as follows:
Ribosome profiling
(Ribo-seq) captures RNA fragments protected by actively translating ribosomes,
providing a genome-wide snapshot of translation with nucleotide resolution.
Polysome profiling separates ribosomes by density on sucrose
gradients, distinguishing mRNAs bound to single ribosomes from those engaged in
multiple-ribosome complexes (polysomes) during active translation.
tRNA sequencing methods quantify tRNA
abundances and their chemical modification. These are crucial regulators of
translation efficiency, often dysregulated in cancer and other diseases.
Advanced techniques such as nano-tRNAseq use nanopore sequencing to directly read native tRNA molecules without reverse transcription, thereby capturing both abundance and modification patterns in a single experiment. Meanwhile, platforms like RiboLace enable active ribosome isolation with significantly less input material (40x less) and faster workflows (3 days vs. 7-10 days) than conventional methods.
1. Biological decisions happen at the translation level
Many
critical biological processes are primarily controlled through translation
rather than transcription. The following are areas where translatomics is
redefining how we see cellular processes:
A) Cancer:
In cancer progression, cells exploit the translation machinery to
produce proteins that promote tumour growth, even when mRNA levels remain
unchanged [2,3]. Cancer cells actively rewire the translation of their mRNAs. A study by Wang
et al.
shows higher translation ratios in lung carcinoma cells than in normal cells
using RNC-seq and Ribo-seq approaches. Moreover, Lian
et al.
utilised the same tools to calculate the elongation velocity index and found
that the translation elongation rate of oncogenes in cancer cells decreased
significantly, thereby ensuring correct folding and malignant function. Even
within the tumour microenvironment, translational regulation shapes immune cell
function and contributes to immune
escape.
B) Neurodegenerative
disease: In neurons, for example, local translation within dendrites and
axons enables spatially restricted protein synthesis that supports synaptic
plasticity, learning, and neuronal signalling, processes that occur far from the
nucleus where transcription takes place [4,5]. Drumond-Bock et al. studied ageing mouse brain using ribosome
profiling and revealed cell type-specific translational changes in dopaminergic
neurons, particularly in mitochondrial and calcium signalling, that likely
explain their vulnerability to neurodegeneration. Separately, Hoorn et al. applied ribosome profiling to motor neurons to
identify translatome-level changes during synapse elimination that were
independent of transcription, and which could be directly targeted to alter
circuit development in vivo. These studies illustrate how translatomics is
uncovering some of the most critical events in neural biology.
C) Cellular
stress: When cells encounter environmental stressors such as heat shock,
hypoxia, or nutrient deprivation, they rapidly reprogram protein production by
selectively regulating translation, often within minutes, while transcriptional
changes typically follow later. This rapid translational control allows cells
to produce pro-survival proteins before new transcripts are produced,
highlighting why studying active protein synthesis is essential for
understanding cellular adaptation and disease mechanisms
[6,7].
2. Identify novel protein-coding sequences hiding in your genome
Ribosome profiling has revealed that the genome encodes far more proteins than
previously recognized. By directly sequencing ribosome-protected mRNA
fragments, this translatomic approach can identify genomic regions that are
actively translated, even when they fall outside conventional gene annotations.
Using this technology, researchers have discovered thousands of previously
unannotated protein-coding sequences, including small open reading frames
(smORFs) that encode functional micropeptides, upstream open reading frames
(uORFs) in the 5′ untranslated regions that regulate downstream translation,
and alternative reading frames that generate additional proteins from known
genes [8,9]. Many smORFs encode microproteins, typically shorter than 100–150 amino acids,
an emerging class of regulatory molecules involved in signaling, metabolism,
and cellular stress responses [10].
Importantly,
large-scale ribosome profiling and proteogenomic studies have demonstrated that
ribosomes frequently occupy regions previously labeled as “non-coding,”
revealing thousands of translated smORFs and novel protein-coding loci across
the human genome
[9,11].
Because these peptides are often small, low-abundance, or embedded within
untranslated regions, they remain largely invisible to traditional gene
annotation pipelines and transcriptomics-based analyses. Translatomics,
therefore, provides a powerful strategy for uncovering this hidden proteome,
expanding our understanding of genome coding potential, and revealing
previously unknown regulatory molecules.
3. Accelerate drug discover and development
Translatomics
is increasingly reshaping the development of RNA therapeutics and mRNA vaccines
by revealing how efficiently therapeutic RNA molecules are converted into
functional proteins inside cells. While the successful delivery of RNA to the
target tissue is essential, the ultimate efficacy of these therapies depends on
the efficient translation of the encoded sequence into the desired protein
product [12]. Techniques such as ribosome profiling and other translatomic approaches enable
direct measurement of the translational output of therapeutic mRNAs, providing
insight into the actual protein synthesis generated from engineered constructs [13]. These methods help researchers optimize coding sequences, codon usage, and
untranslated region (UTR) designs to maximize translation efficiency and
stability of therapeutic transcripts [14].
Translatomics can also reveal unexpected or off-target translation products,
which are critical for safety evaluation in RNA-based therapeutics.
Beyond
RNA medicines, translation profiling is also valuable in small-molecule drug
discovery, particularly for compounds targeting the translation machinery or
ribosome-associated pathways, where measuring changes in ribosome occupancy
helps clarify drug mechanisms of action and identify translational biomarkers
that predict therapeutic response
[15].
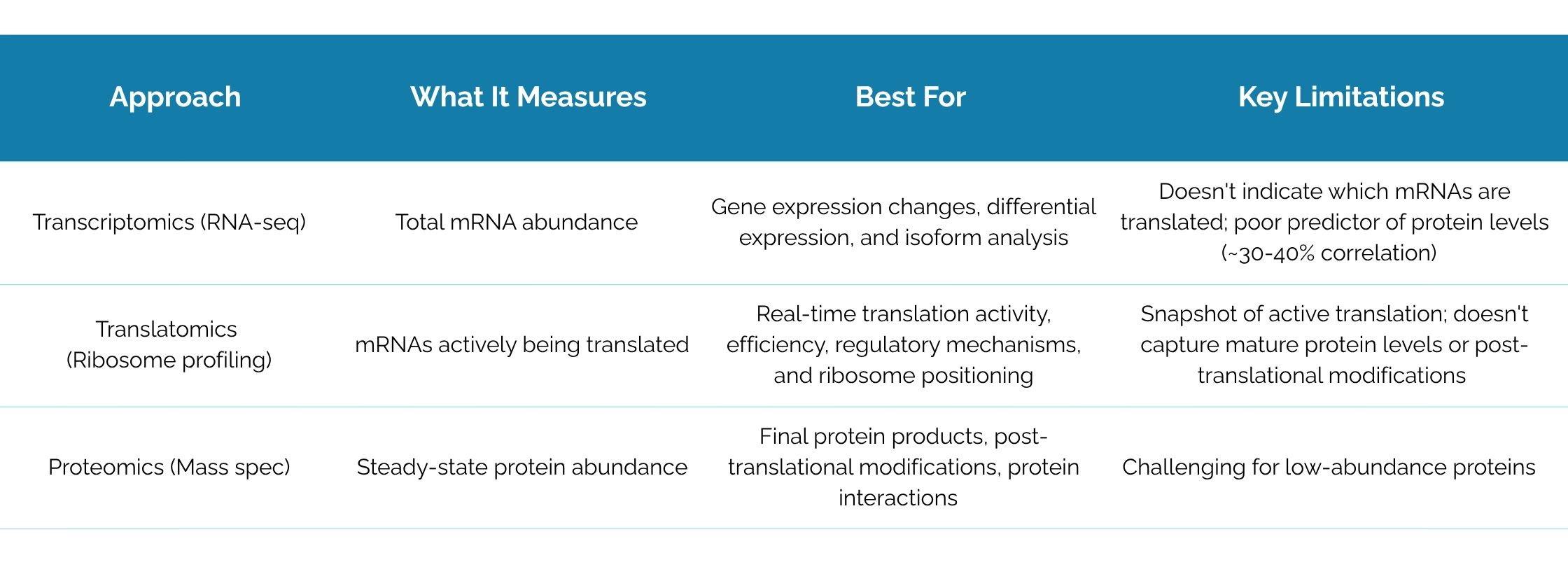
Translatomics vs. other omics: choosing the right approach

Translatomics data: how to analyze and interpret it
Translatomics data
interpretation requires analytical frameworks that are purpose-built for
translation biology. Different approaches produce data sets with unique
characteristics that generic RNA-seq pipelines are not designed to handle.
Proper analysis requires tools that can account for ribosome footprint length
distributions, calculate translation efficiency, model ribosome occupancy, and
integrate tRNA data alongside mRNA measurements.
Immagina’s
bioinformatics Translatome Suite, Martian (mRNAs at ribosomes and tRNAs
included analyzer), is a Python-based platform designed specifically for
translatomic data analysis. MARTIAN was built from the ground up for
translatome analysis, incorporating best practices specific to ribosome
footprint data. MARTIAN guides researchers from raw sequencing reads through
quality control, alignment, quantification, translation-efficiency calculation,
ribosome-occupancy modeling, statistical analysis, visualization, and
functional interpretation.
Learn more
about MARTIAN
The Future of Translatomics
The coming years are
likely to reshape how translation is studied. Over the past decade, ribosome
profiling has revealed the dynamic nature of translation across genomes, but
the next generation of technologies is poised to extend these insights with far
greater spatial, cellular, and analytical resolution.
One of the most exciting frontiers is single-cell translatomics, measuring
ribosome engagement at single-cell resolution. This can help in understanding
cancer, developmental biology, and immunology, where cellular states change
rapidly in response to environmental cues
[16].
Similarly, the integration of spatial transcriptomics with translation
analysis is beginning to illuminate where protein synthesis occurs within
intact tissues. This can allow
researchers to map RNA localization across tissue architecture, and combining
these approaches with measurements of active translation provides a deeper
understanding of how cells regulate protein production in context Sui et al., (2025).
Together, these technological and computational innovations are pushing translatomics
beyond basic research and toward clinical and translational applications.
By directly measuring protein synthesis rather than inferring it from RNA
abundance, translatomics offers a powerful new dimension for biomarker
discovery, therapeutic target identification, and precision medicine, revealing
functional regulatory processes that remain invisible to DNA- or RNA-based
profiling alone [2,3].
Ready to see what your transcriptomics data isn't telling you? Future of Translatomics
Ready to see what your transcriptomics data isn't telling you? Future of Translatomics
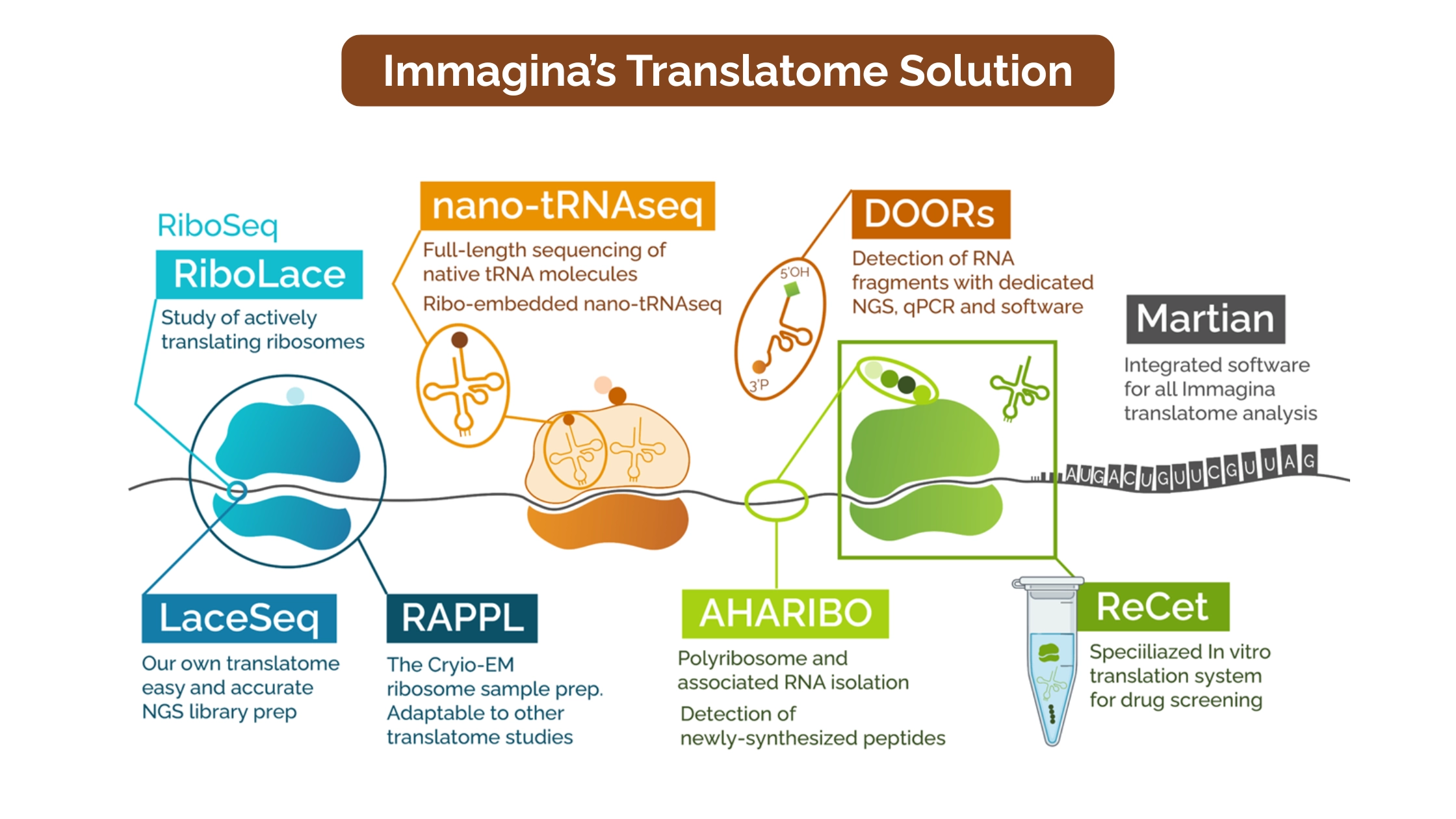
Immagina Biotech's
translatomics platform ranges from gel-free ribosome profiling to sequencing of
native tRNAs to protein detection and analysis. Learn more about our
technologies:
- RiboLace: Gel-free active ribosome profiling requires a minimum starting material of just 35,000 cells, depending on the specimen type, translational activity, and type of experiment.
- nano-tRNAseq: tRNA sequencing with simultaneous detection of tRNA abundance and chemical modifications
- MARTIAN: Dedicated bioinformatics suite for translatome analysis
- DOORs: Detection of 3'-phosphorylated RNA fragments for novel biomarker discovery
Take the next step
Take the next step
Explore Our Technologies – Learn how our platform enables translatomics research
Browse Applications – See how translatomics is advancing disease research
Talk to Our Experts
– Discuss how translatomics can advance your research
Immagina Biotechnology pioneers RNA and ribosome-focused technologies that decode translational landscapes. Our platform supports biotech, pharmaceutical R&D, and academic researchers working on disease mechanisms, drug discovery, and biomarker development. All products are for Research Use Only (RUO).
References and further reading
- Liu, Y., Beyer, A., & Aebersold, R. (2016). On the dependency of cellular protein levels on mRNA abundance. Cell, 165(3), 535-550.
- Ruggero, D. (2013). Translational control in cancer etiology. Cold Spring Harbor perspectives in biology, 5(2), a012336..
- Bhat, M., Robichaud, N., Hulea, L., Sonenberg, N., Pelletier, J., & Topisirovic, I. (2015). Targeting the translation machinery in cancer. Nature reviews Drug discovery, 14(4), 261-278.
- Holt, C. E., & Schuman, E. M. (2013). The central dogma decentralized: new perspectives on RNA function and local translation in neurons. Neuron, 80(3), 648-657.
- Biever, A., Glock, C., Tushev, G., Ciirdaeva, E., Dalmay, T., Langer, J. D., & Schuman, E. M. (2020). Monosomes actively translate synaptic mRNAs in neuronal processes. Science, 367(6477), eaay4991..
- Spriggs, K. A., Bushell, M., & Willis, A. E. (2010). Translational regulation of gene expression during conditions of cell stress. Molecular cell, 40(2), 228-237..
- Liu, B., & Qian, S. B. (2014). Translational reprogramming in cellular stress response. Wiley Interdisciplinary Reviews: RNA, 5(3), 301-305..
- Chothani, S., Ho, L., Schafer, S., & Rackham, O. (2023). Discovering microproteins: making the most of ribosome profiling data. RNA biology, 20(1), 943-954..
- Tong, G., & Martinez, T. F. (2025). Ribosome profiling reveals hidden world of small proteins. Trends in Genetics, 41(2), 101-103..
- Cao, K., Heydary, Y. H., Tong, G., & Martinez, T. F. (2023). Integrated workflow for discovery of microprotein-coding small open reading frames. STAR protocols, 4(4)..
- Kore, H., Okano, S., Datta, K. K., Thorp, J., Periasamy, P., Divate, M., ... & Gowda, H. (2025). Identification of Small Open Reading Frame-encoded Proteins in the Human Genome. Genomics, Proteomics & Bioinformatics, 23(1), qzaf004..
- Pardi, N., Hogan, M. J., Porter, F. W., & Weissman, D. (2018). mRNA vaccines—a new era in vaccinology. Nature reviews Drug discovery, 17(4), 261-279..
- Ingolia, N. T., Ghaemmaghami, S., Newman, J. R., & Weissman, J. S. (2009). Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. science, 324(5924), 218-223..
- Leppek, K., Das, R., & Barna, M. (2018). Functional 5′ UTR mRNA structures in eukaryotic translation regulation and how to find them. Nature Reviews Molecular Cell Biology, 19(3), 158-174.
- Shichino, Yuichi, and Shintaro Iwasaki. "Compounds for selective translational inhibition." Current Opinion in Chemical Biology 69 (2022): 102158.
- Brar, G. A., & Weissman, J. S. (2015). Ribosome profiling reveals the what, when, where, and how of protein synthesis. Nature Reviews Molecular Cell Biology, 16(11), 651-664.