Ribo-tRNAseq
Studying the function of actively used tRNAs in protein translation is fundamental for understanding their role in several cellular processes that can be impaired in various diseases. Immagina’s ribo-tRNAseq leverages our active ribosome pulldown technology, RiboLace, with our tRNA sequencing technology to isolate and sequence ribo-embedded tRNAs. In a single run, our method simultaneously quantifies full-length, native-state tRNAs and detects chemical modifications.
Service overview
Our service ties the pioneering work at the laboratory of Dr. Eva Novoa (Centre for Genomic Regulation (CRG), Barcelona Institute of Science and Technology, Barcelona, Spain with our own active ribosome pulldown innovation. In fact, a recent paper by Monti et al.1 highlights the differences in total tRNA and ribo-tRNA pools upon different stress events, such as translation arrest and viral infection. Ribo-tRNAseq comprises the following steps:
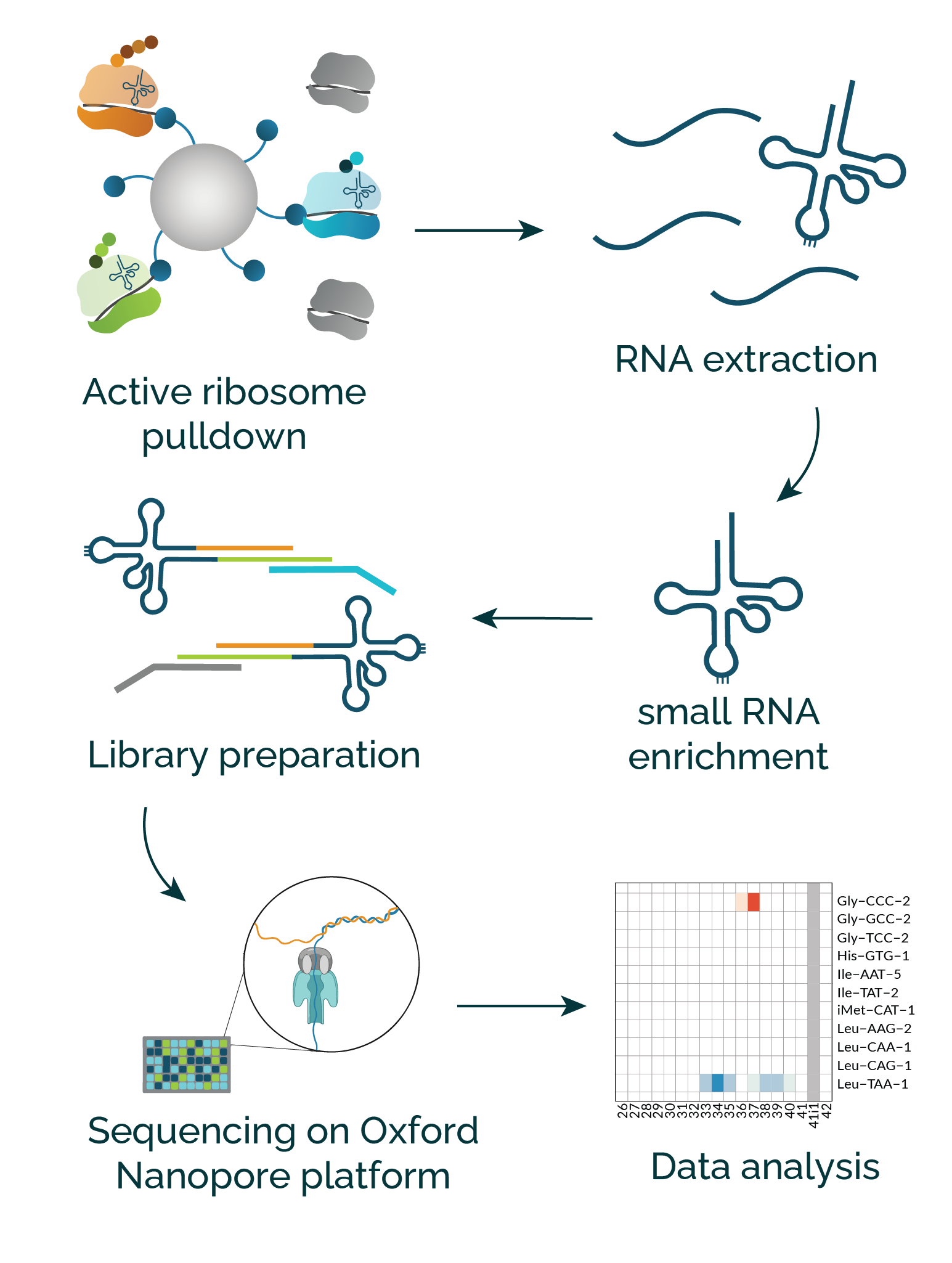
- RiboLace active ribosome pulldown
- Small RNA isolation: enriched from total RNA
- Library preparation: allowing multiplexing up to 6 samples
- Sequencing on Oxford Nanopore platform
- Data analysis
Technical applications
- Uncover abundance and modifications of active tRNAs
- Explore the role of actively used tRNAs in diseases
- Reveal dynamic tRNA usage during translational shifts
- tRNA charging status: upon request
Delivery time: 4 to 6 weeks
Service description and outcomes
- Pre-service consultation to establish your experimental design
- Input material: eukaryotic cell lines (>0.5M) and tissues (>10mg)
- Complete and interactive bioinformatics report
- Access to secure data transfer portal
- Multiplexing up to 6 samples
- High-efficiency data processing

